Understanding the function of PURA in healthy organisms and in the context of PURA Syndrome.
Understanding the function of PURA in healthy organisms and in the context of PURA Syndrome.
The protein PURA (purine-rich element binding protein A) is at the center of scientific investigation due to its involvement in two distinct types of neuronal diseases. In neurodegenerative RNA-repeat expansion disorders like ALS (Amyotrophic lateral sclerosis) and FTD (frontotemporal dementia), PURA has been found to bind to neuronal aggregates. On the other hand, spontaneous de novo mutations of the PURA gene lead to a neurodevelopmental disease known as PURA Syndrome. Despite its significance, our understanding of the molecular pathways affected by PURA in healthy cells and its association with these diseases remains limited.
To bridge this knowledge gap, we teamed up with the groups of Professor Dr. Dierk Niessing at Universität Ulm and Helmholtz Zentrum München. Our primary objective is to shed light on the cellular function of PURA in healthy cells, an endeavor we have already begun to explore in our publication titled “Depletion of the RNA-binding protein PURA triggers changes in posttranscriptional gene regulation and loss of P-bodies” (Molitor & Klostermann et al., Nucleic Acids Research).
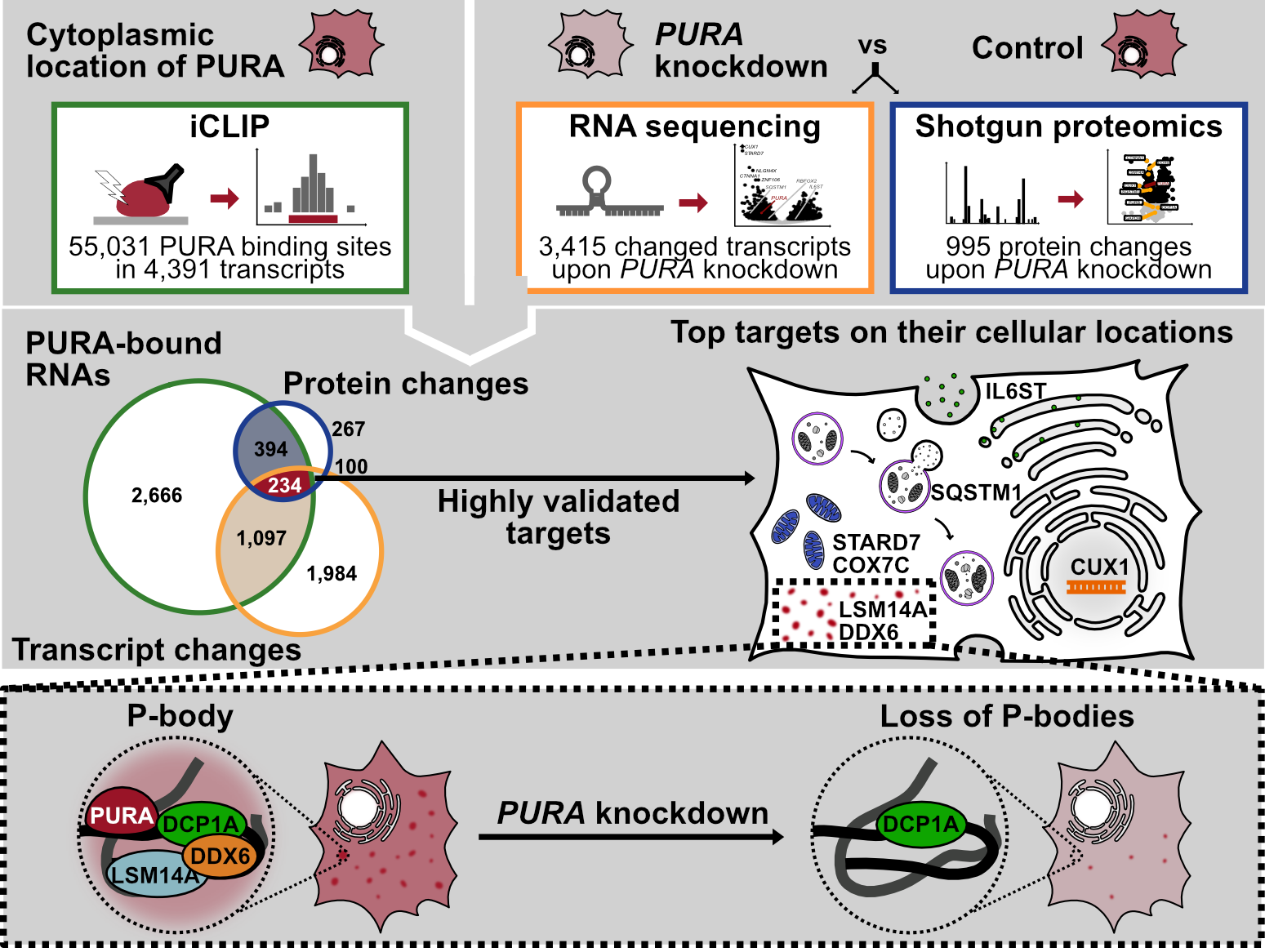
In our publication, we unveil the predominant cytoplasmic localization of PURA, where it interacts with thousands of mRNAs. Remarkably, the depletion of PURA leads to changes in the abundance of numerous transcripts. Based on the proteins encoded by these transcripts, we propose that PURA plays a pivotal role in immune responses, mitochondrial function, autophagy, and processing (P)-body activity. Notably, decreased levels of PURA impact the expression of essential P-body components, LSM14A and DDX6, consequently influencing the formation and composition of this phase-separated RNA processing machinery in human cells.
Building upon these findings, our next step involves expanding our understanding of the effects of PURA syndrome through the utilization of physiological cell lines and diverse disease models.