m6Aboost
The cliProfiler package extension
 {width = “400”, height = “900}
{width = “400”, height = “900}
N6-methyladenosine (m6A) is the most abundant internal modification in mRNA. It impacts many different aspects of an mRNA’s life, e.g. nuclear export, translation, stability, etc.
m6A individual-nucleotide resolution UV crosslinking and immunoprecipitation (miCLIP) and the improved miCLIP2 are m6A antibody-based methods that allow the transcriptome-wide mapping of m6A sites at a single-nucleotide resolution (Körtel et al. 2021)(Linder et al. 2015). In brief, UV crosslinking of the m6A antibody to the modified RNA leads to truncation of reverse transcription or C-to-T transitions in the case of readthrough. However, due to the limited specificity and high cross-reactivity of the m6A antibodies, the miCLIP data comprise a high background signal, which hampers the reliable identification of m6A sites from the data.
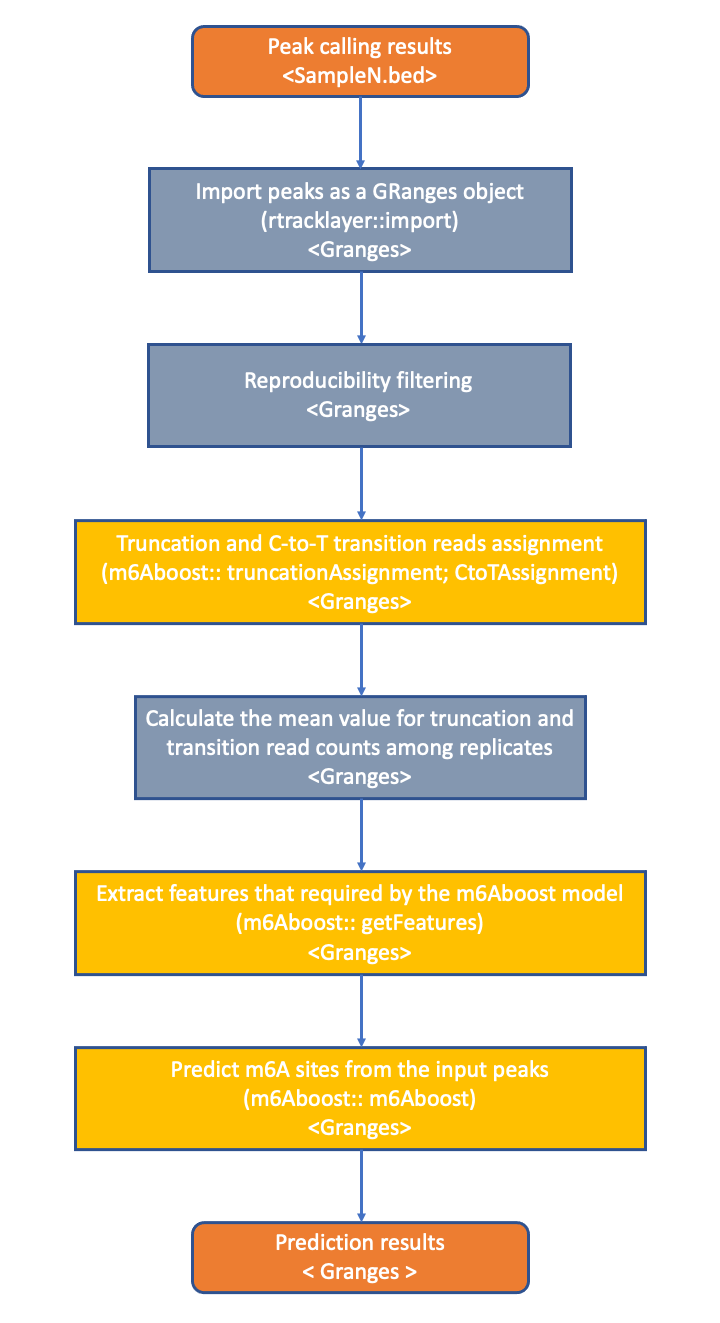
For accurately detecting m6A sites, we implemented an AdaBoost-based machine learning model (m6Aboost) for classifying the miCLIP2 peaks into m6A sites and background signals (Körtel et al. 2021). The model was trained on high-confidence m6A sites that were obtained by comparing wildtype and Mettl3 knockout mouse embryonic stem cells (mESC) lacking the major methyltransferase Mettl3. For classification, the m6Aboost model uses a series of features, including the experimental miCLIP2 signal (truncation events and C-to-T transitions) as well as the transcript region (5’UTR, CDS, 3’UTR) and the nucleotide sequence in a 21-nt window around the miCLIP2 peak.
The m6Aboost package is available at https://bioconductor.org and can be access with the following link: https://bioconductor.org/packages/release/bioc/html/m6Aboost.html.